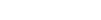
Perelman School of Medicine researchers at the University of Pennsylvania are developing a new type of positron emission tomography (PET) scan for imaging the brains of patients with neurodegenerative disorders, including Parkinson’s disease.
Penn Medicine researchers will work with scientists from the Washington University-St Louis, the Universities of Pittsburgh and California-San Francisco and Yale University.

Discover B2B Marketing That Performs
Combine business intelligence and editorial excellence to reach engaged professionals across 36 leading media platforms.
The alliance, Center Without Walls, secured a $20m National Institute of Neurological Disorders and Stroke (NINDS) grant to support the project over five years.
Center Without Walls principal investigator Robert Mach said: “At the end of five years, we hope to have a radioactive tracer that will be able to detect Parkinson’s early on and provide detailed information about the disease’s progression, which is critical for discovering and testing new treatments.”
Parkinson’s disease, which currently lacks a diagnostic test, could remain undetected or be misdiagnosed until its symptoms become severe.
Treatments could become less effective with disease progression and an imaging biomarker or indicator is expected to allow early diagnosis, accelerating clinical trials of drugs.
US Tariffs are shifting - will you react or anticipate?
Don’t let policy changes catch you off guard. Stay proactive with real-time data and expert analysis.
By GlobalDataDuring a PET scan, a radioactive drug or tracer binds to proteins or sugars to highlight areas of the body with higher chemical activity levels, which indicate disease.
As part of the project, researchers are working to detect Parkinson’s and other neurodegenerative diseases driven by proteinopathies, which form when some proteins ‘misfold’ and become structurally abnormal.
The team plans to create a radiotracer that will bind to the alpha-synuclein protein in the brain for imaging of Parkinson’s and multiple system atrophy.
Also, they will develop another radiotracer, which will bind to the 4R tau protein, to image frontotemporal degeneration and progressive supranuclear palsy.
To develop the radiotracers, the team will leverage a computational technology that screens for molecules, synthesises them and interprets binding data depending on crosslinking.